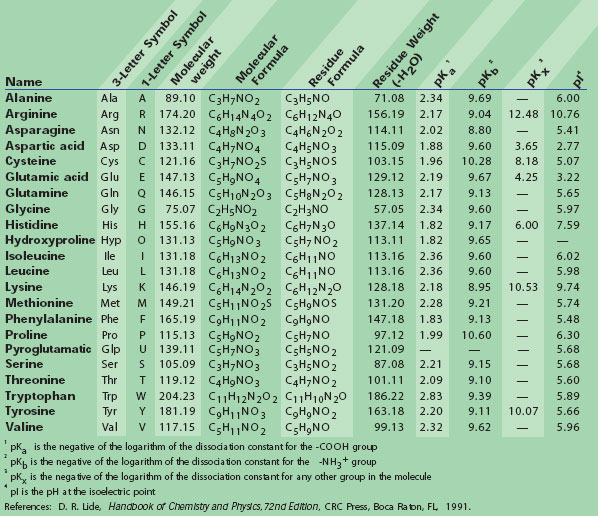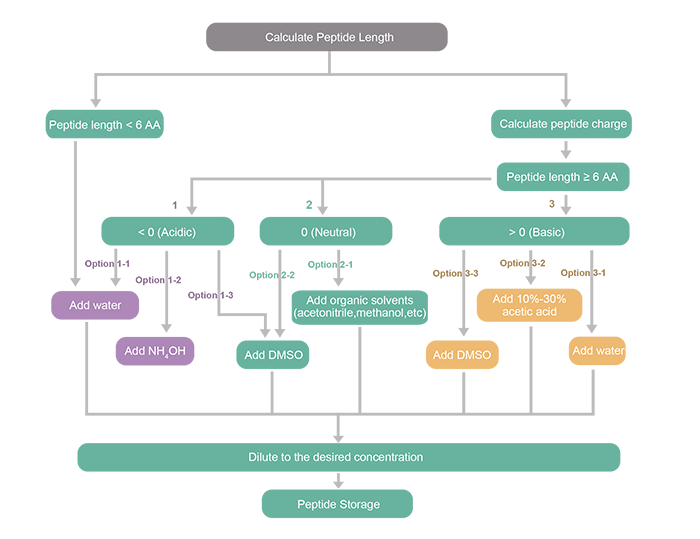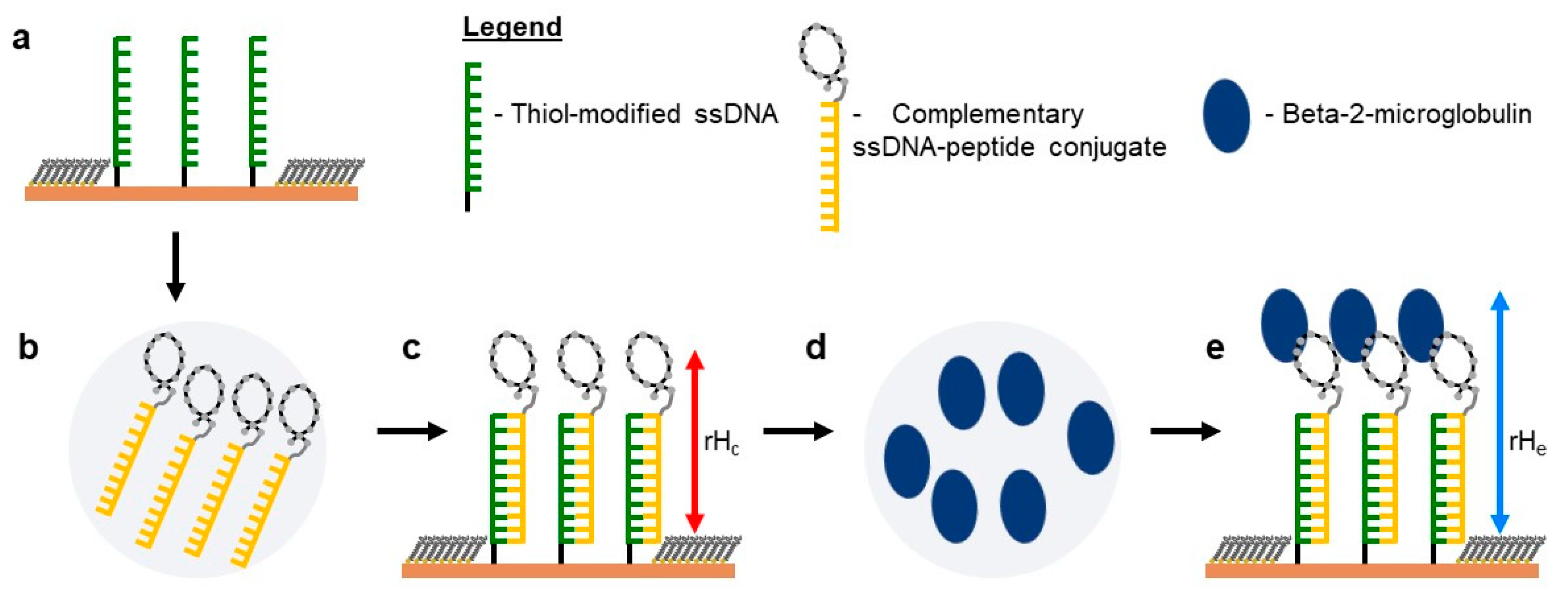
IJMS | Free Full-Text | Computational Evolution of Beta-2-Microglobulin Binding Peptides for Nanopatterned Surface Sensors

pIChemiSt ─ Free Tool for the Calculation of Isoelectric Points of Modified Peptides | Journal of Chemical Information and Modeling

Solubility test of NSP and NSP r. (A) Peptide models computed by the... | Download Scientific Diagram

Melting Properties of Peptides and Their Solubility in Water. Part 2: Di- and Tripeptides Based on Glycine, Alanine, Leucine, Proline, and Serine | Industrial & Engineering Chemistry Research

Sequence-based prediction of the intrinsic solubility of peptides containing non-natural amino acids | Nature Communications

pIChemiSt ─ Free Tool for the Calculation of Isoelectric Points of Modified Peptides | Journal of Chemical Information and Modeling
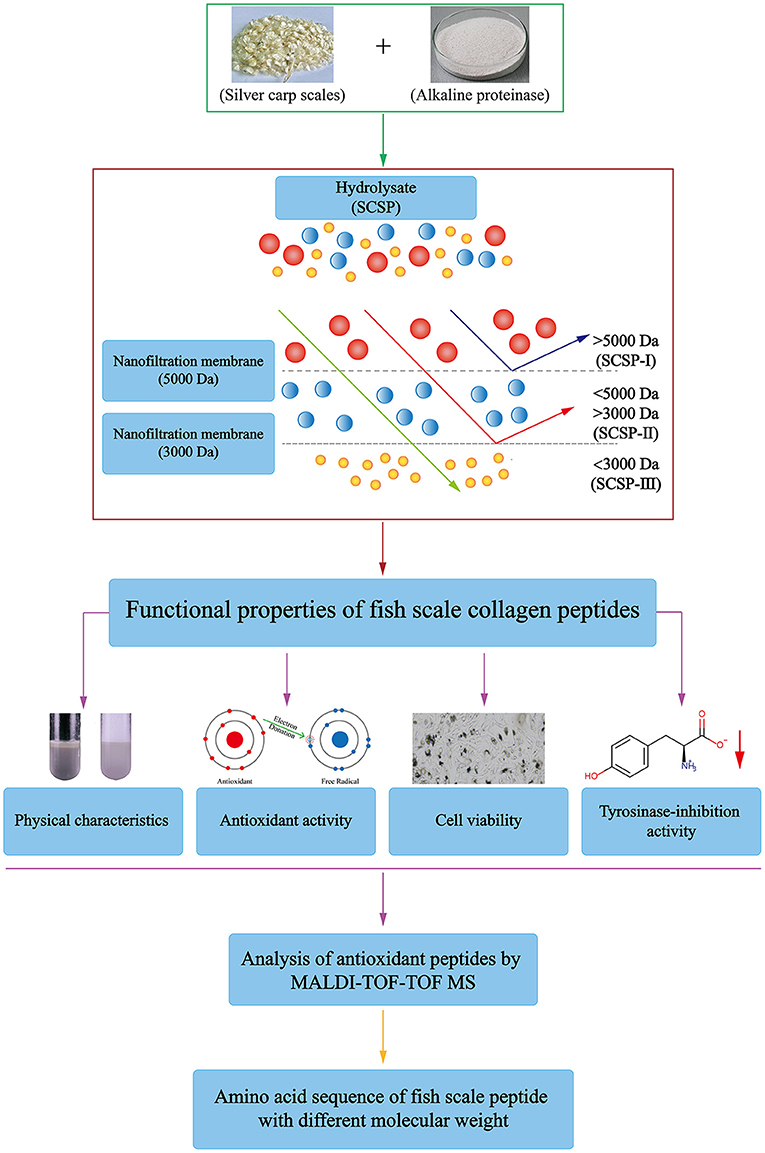
Frontiers | Physicochemical Properties and Biological Activities of Silver Carp Scale Peptide and Its Nanofiltration Fractions

A comparative study of synthetic winged peptides for absolute protein quantification | Scientific Reports



.png)